Main Menu (Mobile)- Block
- Our Research
-
Support Teams
- Overview
- Anatomy and Histology
- Cryo-Electron Microscopy
- Electron Microscopy
- Flow Cytometry
- Fly Facility
- Gene Targeting and Transgenics
- Immortalized Cell Line Culture
- Integrative Imaging
- Janelia Experimental Technology
- Media Prep
- Molecular Genomics
- Primary & iPS Cell Culture
- Project Pipeline Support
- Project Technical Resources
- Quantitative Genomics
- Scientific Computing Software
- Scientific Computing Systems
- Viral Tools
- Vivarium
- Open Science
- You + Janelia
- About Us
Main Menu - Block
- Overview
- Anatomy and Histology
- Cryo-Electron Microscopy
- Electron Microscopy
- Flow Cytometry
- Fly Facility
- Gene Targeting and Transgenics
- Immortalized Cell Line Culture
- Integrative Imaging
- Janelia Experimental Technology
- Media Prep
- Molecular Genomics
- Primary & iPS Cell Culture
- Project Pipeline Support
- Project Technical Resources
- Quantitative Genomics
- Scientific Computing Software
- Scientific Computing Systems
- Viral Tools
- Vivarium
Publications
Featured

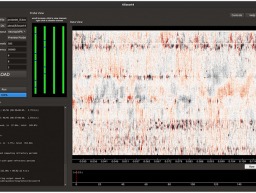




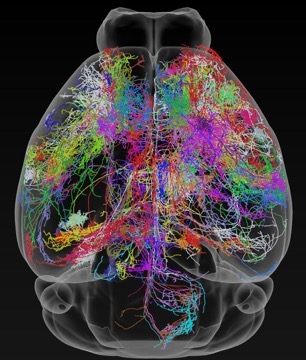
A vast neural tracing effort by a team of Janelia scientists has upped the number of fully-traced neurons in the mouse brain by a factor of 10. Researchers can now download and browse the data in three dimensions.
Use NeuronBrowserThese split-GAL4 driver Drosophila lines allow the generation of cell-type specific gene expression. They are freely available through the Bloomington Stockcenter through a large coordinated effort from Janelia Drosophila Resources and the National Institutes of Health.
Learn MoreThese bright and photostable fluorescent labels enable imaging and tracking of individual molecules inside living cells.
Learn MoreThe GCaMP6 family of GECIs is a collection of green fluorescent indicator proteins that facilitate the measurement of synaptic calcium signals, enabling reliable detection of neuronal responses in vivo. The next-generation GCaMP6s, GCaMP6m, and GCaMP6f sensors offer increased sensitivity (single action potential detection in vivo) and improved kinetics.
Learn MoreThis photoconvertible protein enables imaging of the calcium activity history of large areas and populations of cells. It is based on EosFP, a fluorescent protein that shows emission changes from green to red upon irradiation with UV light. It is not restricted to the field of view of a microscope, as during real-time calcium imaging.
Learn MoreThis microscope uses a Bessel beam to illuminate a sample with light sheets that are sufficiently thin to achieve isotropic resolution high-speed 3D fluorescent imaging. The technology can visualize cellular processes in spatiotemporal detail not attainable with other fluorescent microscopy techniques.
Learn More